Lab
Biology Bldg. 425
van Kessel Lab website
Research
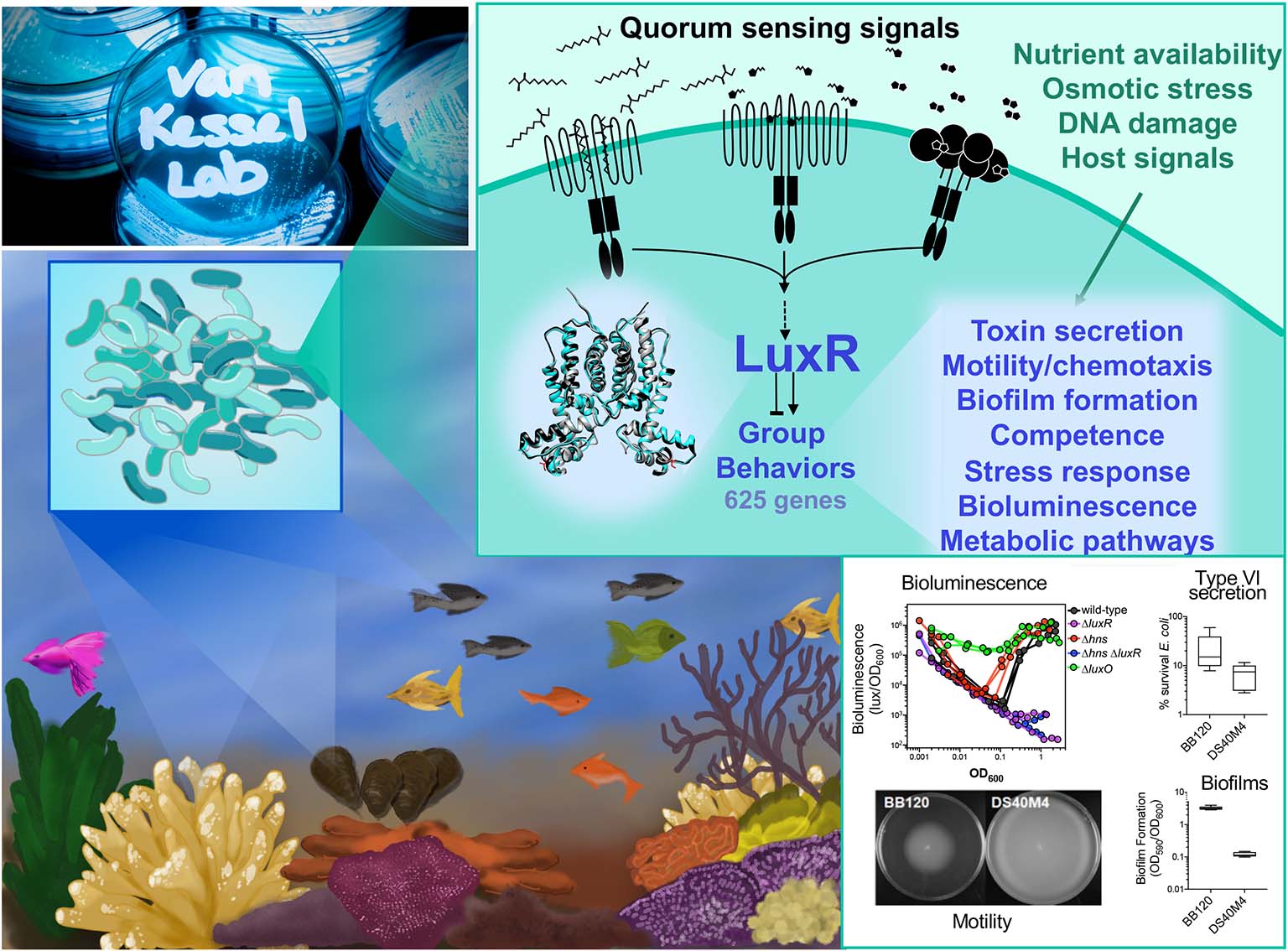
The van Kessel lab studies communication and behavior in pathogenic Vibrio bacteria. Bacteria use the cell-cell communication process called quorum sensing to sense and respond to changes in the number and type of cells in their surrounding community. Through quorum sensing, bacterial cells synchronously alter gene expression to change modes of behavior. We are interested in the molecular, genetic, and biochemical mechanisms that coordinate this developmental switch. Our lab uses a unique multi-disciplinary approach to study quorum sensing in vibrios, including microbial genetics, biochemistry, molecular biology, chemistry, structural biology, transcriptomics, and bioinformatics.
The focus of our research is on marine Vibrio bacteria, which are pathogens of fish, shellfish, coral, and mammals, including humans. Rising ocean temperatures have driven increased Vibrio infections in natural marine habitats including coral reefs, fish and shellfish aquaculture, and in humans due to consumption of contaminated seafood. Many of the processes controlled by quorum sensing influence cell growth and development, including pathogenesis, symbiosis, and responses to nutrient stress. Our research aims to gain a mechanistic understanding of how cell-cell communication directly impacts bacteria in their environmental niches.
Research Areas
Genomics and Bioinformatics
Microbial Cell Biology and Environmental Responses
Microbial Interactions and Pathogenesis


 The College of Arts
The College of Arts